Imagine if we could “grow” a digital replica of a living cell inside a computer and observe how it responds under various conditions. A recently published review has outlined the contours of this possibility, centering on the cutting-edge concept of Virtual Cells, and depicting an emerging field that is moving from academic research to biomedical applications. Virtual cells are not a fantasy, but a tangible technology integrating multi-omics data and computational modeling, with ongoing exploratory applications in drug development, precision medicine and other fields. For assisted reproduction, a field that relies heavily on empirical judgment yet has long been trapped by a “bottleneck in success rates”, this technological direction deserves attention.

Understanding Virtual Cells
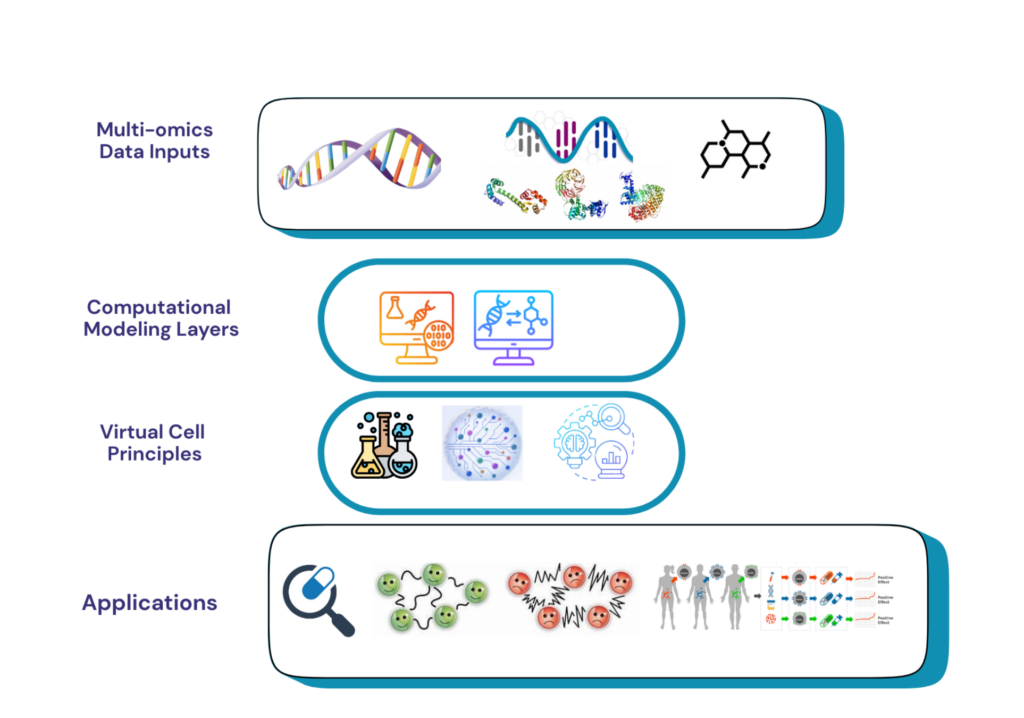
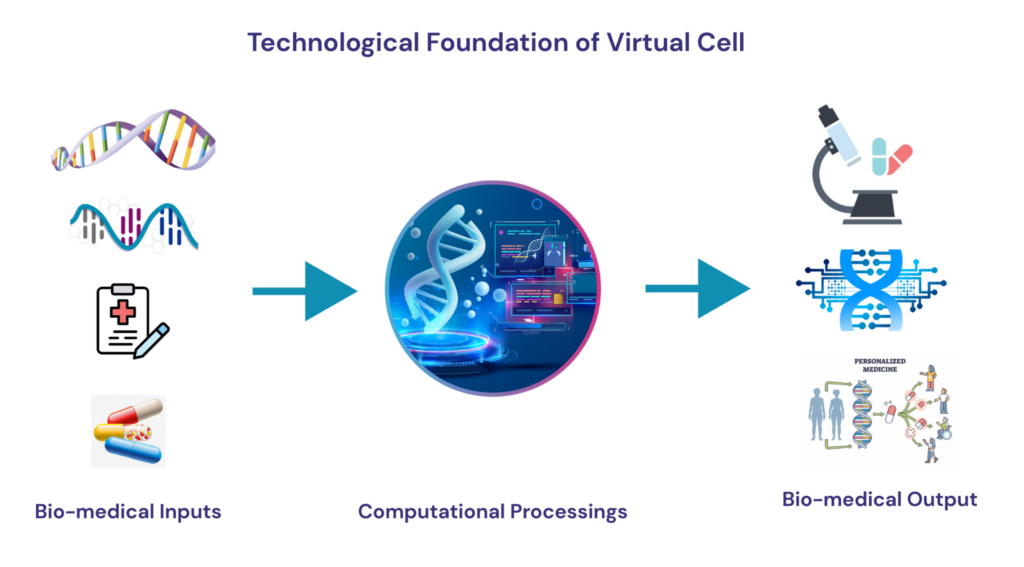
In simple terms, a virtual cell is a “digital clone” of a cell. It does not simply replicate the appearance of a cell, but reconstructs the internal operational logic — how genes are regulated, how proteins interact, how signals are transduced, and how energy flows. It can be compared to building a precise architectural model: instead of only looking at the finished product, we clarify the specifications and connections of every beam, column and pipeline, so that the model can be used to simulate the performance of a real building under various conditions. Building such a digital model first requires using various omics technologies to portray the molecular state of cells from different perspectives; then integrating these data into a unified computational framework to reconstruct gene regulatory networks, protein-protein interaction networks and metabolic pathway maps; finally, introducing various perturbations into the model to observe cellular responses. At present, this field has moved from proof of concept to practical application exploration. Although it is still far from the ideal complete cell simulation, the technical pathway is clear and feasible.
Virtual Cells and IVF
The daily work of assisted reproduction is essentially dealing with “uncertainty”: every step from oocyte to embryo is full of unknowns. Existing technologies — morphological observation, time-lapse imaging, multi-omics testing — each provide fragmented information, but how to integrate these fragments into a complete understanding has always been a core problem plaguing this field. The core value of virtual cells lies precisely in this — it is not a new technology, but a framework for integrating multi-dimensional information, simulating cell behavior and understanding developmental mechanisms.
The integration dilemma of developmental potential assessment is the first typical scenario. The developmental potential of an oocyte is not determined by a single indicator, but jointly by multiple factors such as gene expression regulation, protein translation, mitochondrial function and signal pathway transduction. Morphological indicators reflect the final result rather than the process, and multi-omics studies are limited by the difficulty of sample acquisition. Existing artificial intelligence algorithms have begun to learn patterns from multi-omics data, but they are essentially still “input-output” statistical mappings lacking in-depth understanding of mechanisms. What virtual cells attempt to do is integrate these scattered molecular events into a unified dynamic model. The model can not only judge “how is the developmental potential of this oocyte”, but also explain “why its developmental potential may be at this level” — which is crucial for understanding the direction of intervention.
The “black box” dilemma in understanding fertilization is the second typical scenario. Fertilization is a highly complex sequence of molecular events: sperm capacitation, acrosome reaction, sperm-egg fusion, pronuclear formation… Each link involves the spatiotemporal coordination of dozens of molecules. At present, the understanding of fertilization mainly comes from observations of final results and inferences from animal models — it is almost impossible to fully track the dynamic process of fertilization in living cells. Computational modeling offers a new possibility to break this dilemma. When the virtual cell model is sufficiently refined, researchers may be able to simulate the molecular dynamics of fertilization in computers, explore fertilization trajectories under different conditions, and identify key nodes that may lead to fertilization failure.
The leap from observation to understanding is the profound value that virtual cells bring to assisted reproduction. Embryologists observe embryo morphology and record developmental trajectories day in and day out, but what exactly do these observations mean — why is the cleavage speed of this embryo slow? Why do fragments suddenly appear in that embryo at a certain time point? Traditionally, these questions can often only stay at the descriptive level, making it difficult to delve into molecular mechanisms. Virtual cells provide a possibility to connect “observed phenomena” with “mechanistic understanding”. The value of the model lies not only in predicting “this embryo may be abnormal”, but also in explaining “what molecular events caused it to be abnormal” — and the latter is the key to truly guiding intervention decisions.
Clinical Value and Prospects
The clinical value of virtual cell technology for assisted reproduction is reflected in at least three levels. Deepening of evaluation dimensions: when digital models can simulate intracellular molecular dynamics, embryo evaluation will no longer be limited to morphological and kinetic parameters, but may extend to deep features such as metabolic status, signaling pathway activity, and stability of gene regulatory networks. Personalization of decision support: building exclusive digital cell models for specific patients and simulating the expected outcomes of different treatment plans in the model. Once this “compute first, act later” model matures, it will greatly improve the adequacy of information for clinical decision-making. Improvement of research efficiency: virtual cells can serve as a virtual experimental platform for mechanistic research, making precious real samples used where they matter most.
Of course, there is still a long way to go from vision to reality. The scale and quality of data accumulation, the accuracy verification of computational models, and the integration with existing workflows — there are practical challenges in every link. However, this technology has the potential of “self-evolution”: with the continuous accumulation of clinical data and the continuous advancement of model verification, the prediction accuracy of virtual cells will gradually improve, forming a virtuous cycle of “more accurate with more use”. For assisted reproduction, a field long trapped by the “glass ceiling of success rates”, this may be a new path worthy of serious attention.
Conclusion
There is still a long way to go for virtual cells to move from academic concepts to clinical applications. But it provides a new thinking framework for reproductive medicine: when observational methods reach a bottleneck, perhaps we can change a dimension to find answers. No matter how the final technological form evolves, the continuous exploration of embryonic developmental mechanisms will eventually promote the progress of the entire field. When that day comes, virtual cells may, like genome sequencing today, change from cutting-edge research tools to routine clinical tools, which will be another leap forward in reproductive medicine.
